Understanding Bulk Multiomics: A Technical Introduction
As multiomics becomes increasingly central to biological research and therapeutic development, scientists are seeking practical, scalable approaches to generate integrated datasets across molecular modalities. While single-cell multiomics and spatial multiomics receive significant attention, bulk multiomics remains a powerful and accessible strategy for profiling transcriptomic, proteomic, and other molecular features across entire samples or tissue sections.
This article offers a technical overview of bulk multiomics: what it is, how it compares with single-cell approaches, where it fits in modern research workflows, and how emerging tools are making it easier to adopt at scale.
On this page:
Multiomics Enables a More Complete View of Biology
Traditional transcriptomic or proteomic methods each offer a partial view of cellular state. However, mRNA abundance does not always correlate with protein levels or activity, and proteins are subject to post-translational modifications that impact function. Integrating RNA and protein data from the same sample can:
- Validate findings across molecular layers
- Reveal regulatory checkpoints and disconnects
- Support functional interpretation of biological states
- Improve reproducibility and biological relevance
Together, these insights support a systems-level understanding of pathways, cell states, and responses, which are essential for advancing research in fields such as cancer, immunology, and regenerative medicine.
Bulk Multiomics Versus Single-Cell Multiomics
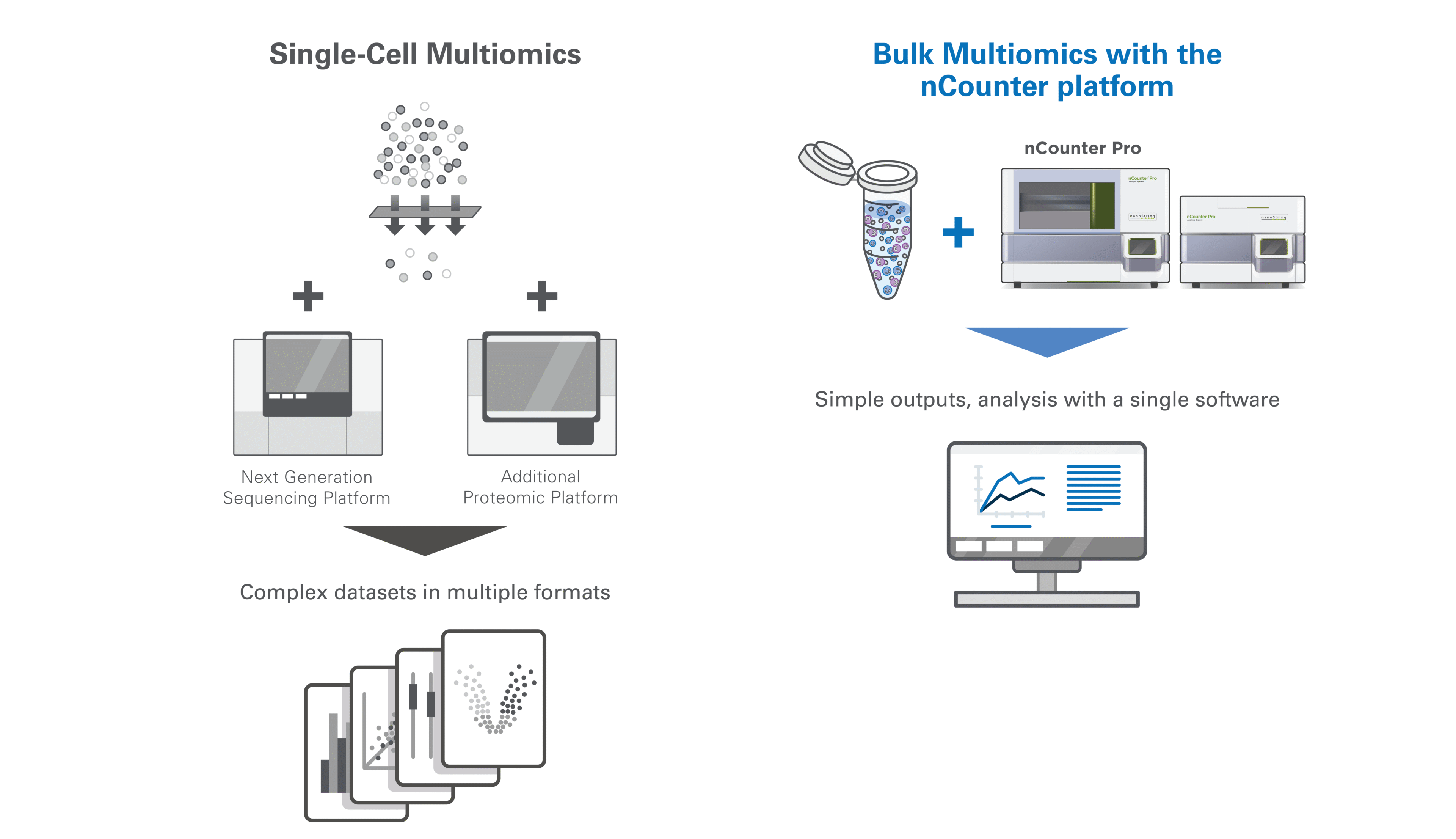
The key distinction between bulk and single-cell multiomics lies in resolution. Single-cell methods measure RNA and protein expression within individual cells, enabling high-resolution insights into heterogeneity, rare populations, and cell-cell interactions. However, they come with tradeoffs: sparse data, complex sample preparation, and limited scalability.
Bulk multiomics, by contrast, aggregates signals across cell populations. This enables simpler workflows, robust signal quantification, and greater scalability for studies involving large cohorts or longitudinal timepoints. While single-cell resolution is lost, bulk approaches offer advantages in stability, reproducibility, and interpretability.
Bulk Multiomics Versus Single-Cell Multiomics
Spatial multiomics technologies extend molecular profiling by preserving tissue architecture and spatial context. These methods allow researchers to localize RNA and protein within intact tissue sections, enabling a deeper understanding of cellular microenvironments and interactions.
Platforms like the GeoMx® Digital Spatial Profiler and the CosMx® Spatial Molecular Imager are leaders in spatial multiomics. The GeoMx platform enables the highest-plex profiling of RNA and protein in FFPE samples with regional resolution, making it ideal for discovery applications. The CosMx platform enables high fidelity spatial multiomics with single-cell resolution across entire tissue sections.
Compared to bulk and single-cell methods, spatial multiomics provides:
- Spatial localization of molecular signals
- Compatibility with complex tissues and clinical samples
- Insights into cell-cell interactions and tissue architecture
While spatial approaches offer unique advantages, they can be technically demanding and resource-intensive. For many applications, combining bulk, single-cell, and spatial data is the ideal way to uncover a complete and comprehensive biological picture.
When to Use Bulk Multiomics
Bulk multiomics is well-suited for applications in oncology, immunology, and translational research, particularly when:
- Sample material is limited, and maximizing data from a single input is critical
- Study designs require large cohorts, replicates, or time-course experiments
- Averaged signals are sufficient to capture biological effects
Combining transcript and protein measurements enables more nuanced interpretation of regulatory events, pathway dynamics, and disease mechanisms. Hybrid strategies that integrate bulk and single-cell data are also increasingly common, especially when paired with machine learning models.
Applications in Life Science Research
1. Biomarker and Signature Development
Multiomic profiling enables development of more robust molecular signatures by combining mRNA expression with protein abundance data. For example:
- In oncology, immunotherapy response models have been improved by combining transcript-based immune profiles with protein-level expression of checkpoint molecules like PD-L1.
- In inflammatory disease research, co-detection of cytokine transcripts and their soluble receptors offers a more accurate view of pathway activation.
2. Target Identification and Drug Discovery
Multiomics supports drug development by linking gene expression to protein function:
- Neurodegeneration studies have mapped changes from transcriptional dysregulation to synaptic protein loss.
- Early-stage screening uses multiomic profiles to confirm target expression and functional relevance in disease models.
3. Regulatory Insights and Hypothesis Generation
Discrepancies between RNA and protein measurements can uncover post-transcriptional regulation, degradation, or feedback mechanisms:
- In autoimmune diseases, elevated cytokine transcripts alongside suppressed proteins have revealed novel control points.
- In stem cell differentiation, tracking RNA and surface proteins maps lineage progression with high precision.
Introducing Scalable, Accessible Bulk Multiomics
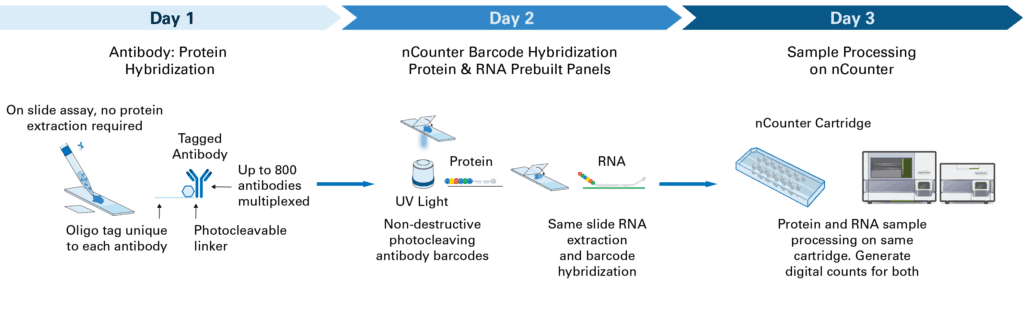
Until recently, technical limitations often required researchers to compromise between data types, input requirements, or throughput. Our nCounter® multiomics technology closes that gap, allowing scalable quantification of RNA and protein targets from a variety of sample types, with the same robust and reliable performance that the nCounter platform has achieved for nearly two decades.

This new multiomic approach integrates direct digital detection of both RNA and protein analytes, maintaining high specificity and accurate quantification. Combined with the established simplicity and reproducibility of the nCounter Analysis System, it offers a practical solution for researchers seeking to scale bulk multiomics to larger cohorts or tissue studies.
We also offer a Technology Access Program (TAP), providing scientists early access to this platform for pilot studies or exploratory projects—ideal for teams looking to evaluate multiomic workflows without investing in infrastructure upfront.
Final Thoughts
Bulk multiomics provides a powerful complement to single-cell and spatial approaches. By enabling multiplexed RNA and protein profiling across tissue sections or cell populations, it delivers the integrated molecular data needed for cohort-scale discovery, biomarker validation, and systems-level modeling. As new technologies lower barriers to adoption, we expect bulk multiomics to take on a larger role in translational and preclinical research pipelines.
References
Stay informed about bulk multiomics advances
Join our mailing list for the latest updates on nCounter multiomics technology.