Accurate Cell Segmentation: The Critical Foundation of Spatial Biology
Authors: Kalyan Chavda, Lidan Wu, Aster Wardhani
Spatial biology has entered the era of single-cell and subcellular resolution and generating high-dimensional molecular data is no longer the challenge. Interpreting it accurately is. At the core of this is cell segmentation: the process of delineating molecular data into functional cellular units (“cells”) by defining true cell boundaries. This has emerged as a critical determinant of data quality and biological insights that can be derived.
Single-cell spatial imaging technologies now enable researchers to map gene and protein expression directly within intact tissue. However, accurately distinguishing cell boundaries remains a major challenge and can significantly impact downstream analysis (Petukhov et al., Nature Biotechnology, 2022; Mitchel et al., Nature Genetics, 2026). In this context, cell segmentation is not simply a preprocessing step—it is the foundation upon which all downstream insights are built.
- Why Cell Segmentation Matters
- The Challenge of Segmenting Real World Tissue Sections
- Advancing Cell Segmentation with Multimodal and AI-Driven Approaches
- Best-in-Class Cell Segmentation with CosMx® Spatial Molecular Imager (SMI)
- From Segmentation to Biological Insight
- Setting a New Standard in Spatial Biology
Why Cell Segmentation Matters
Accurate cell segmentation enables a deeper understanding of biology at multiple levels:
- Understanding Cellular Interactions: Cell segmentation delineates biological data into individual cells, enabling study of individual cells and interactions across cells and pathological conditions.
- Analyzing Biological Features: Cell segmentation empowers scientists to analyze essential biological features like cell count, type, shape, and more. This analysis provides insights into how these features change over time and in response to different conditions.
- Capturing Biologically Relevant Information: The morphology of cells reflects their physiological state, and effective cell segmentation captures biologically relevant morphological information.
- Accelerating Drug Discovery: Insights gained from cell segmentation can drive drug discovery, diagnostics, and various other critical areas in biology, pharmacology, and personalized medicine.
Importantly, spatial transcriptomics uniquely preserves the relationship between gene expression and spatial context, allowing researchers to study how cellular function is shaped by tissue architecture (Liu et al., 2022). In summary, cell segmentation is a cornerstone of single-cell spatial research, playing a fundamental role in understanding health and disease.
The Challenge of Segmenting Real World Tissue Sections
Despite its importance, accurate cell segmentation remains challenging, particularly in complex biological tissues. Accurate cell segmentation can be a daunting task due to several factors:
- Heterogeneous shapes: Cells come in diverse and dynamic shapes, making it nearly impossible to define mathematical shape models
- Variation in cell size and morphology: unlike nuclei, which are typically similar-sized structures, the cytoplasm exhibits significant variations in shape and size
- Weak boundary gradients, making it difficult to distinguish adjacent cells
- Dataset-specific variability, including differences in staining, imaging modalities, artifacts in image capture, and tissue types
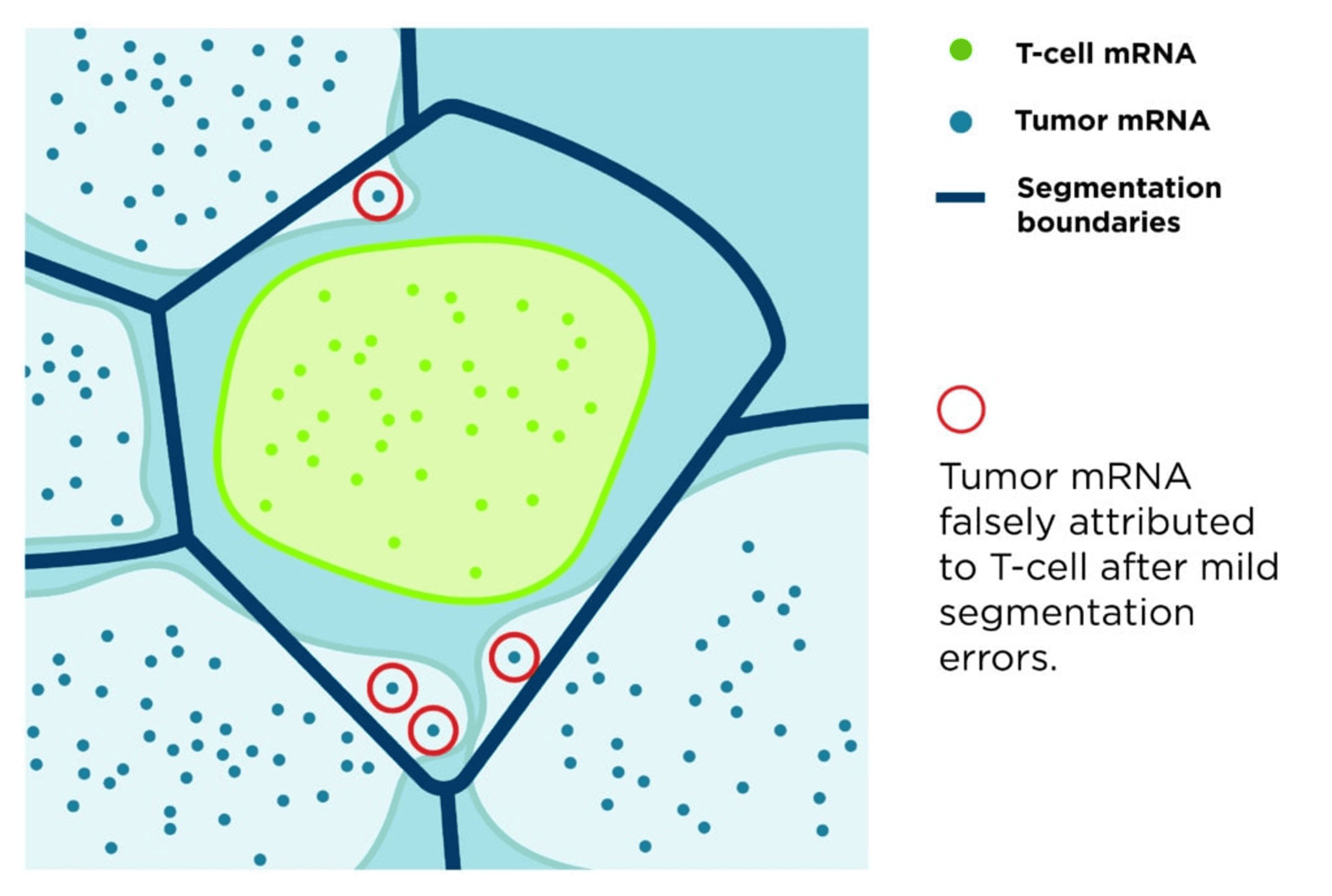
Many traditional approaches rely on nuclear expansion methods, where cell boundaries are approximated by extending outward from the nucleus. While computationally convenient, these approaches frequently result in systematic misassignment of transcripts, particularly in complex or dispersed cell types.
This limitation was well recognized: traditional segmentation approaches often approximate cell boundaries using nuclear stains, reducing accuracy and introducing artifacts (Petukhov et al., Nature Biotechnology, 2022). These errors propagate through downstream analyses, distorting cell identity, masking biological heterogeneity, and compromising biological interpretability (Mitchel et al., Nature Genetics, 2026).
Advancing Cell Segmentation with Multimodal and AI-Driven Approaches
To overcome these challenges, the field is rapidly evolving toward multimodal and machine learning–based segmentation strategies. Recent advances demonstrate that combining RNA spatial information with nuclear and membrane markers significantly improves segmentation accuracy and transcript assignment (Defard et al., Genome Biology, 2026). These approaches move beyond purely image-based models, leveraging the joint distribution of molecular and morphological data.
Deep learning frameworks such as Cellpose and related models further enhance segmentation by learning from diverse tissue types and imaging conditions, improving robustness across complex biological samples (Stringer et al., Nature Methods, 2021; Jin et al., Nature Communications, 2025).
However, even advanced models face limitations. They may struggle with trade-offs between capturing elongated morphologies and resolving densely packed cells, leading to over- or under-segmentation in challenging regions. This underscores that segmentation remains an open and evolving problem.
Best-in-Class Cell Segmentation with CosMx® Spatial Molecular Imager (SMI)
From the very beginning, the CosMx® SMI employed a fundamentally robust approach: one that integrates multimodal data and advanced machine learning to define true cellular boundaries. At the core of CosMx segmentation is an enhanced implementation of the CellPose algorithm (Stringer et al., Nature Methods, 2021), trained and validated on millions of cells across diverse tissue types. Rather than inferring cell boundaries, CosMx precisely defines them by combining:
- Nuclear markers
- Cell membrane and cytoplasm protein signals
- Spatial RNA information
This multimodal strategy enables precise delineation of complex cell morphologies, significantly improving accuracy over methods based solely on nuclear expansion. Recent advances from Bruker Spatial Biology further extend this capability with the 3D AI segmentation models implemented in the AtoMx® Spatial Informatics Platform. By reconstructing cells across multiple focal planes, these models resolve overlapping cells and improve single-cell boundary definition in dense tissues, overcoming long-standing limitations of traditional segmentation approaches. Finally, the segmentation models are purpose-trained using machine learning on diverse tissue types, enabling robust performance across complex samples such as brain tissues.
From Segmentation to Biological Insight
The impact of segmentation accuracy is profound. Cell segmentation errors can lead to incorrect gene assignment, introducing artifacts in clustering, differential expression, and cell-cell interactions. In contrast, precise segmentation preserves the integrity of spatial data, enabling researchers to confidently interpret cellular identity, function, and interaction within tissue context.
By minimizing segmentation errors and accurately capturing complex cell shapes, CosMx SMI enables:
- Higher fidelity single-cell data
- Improved detection of rare and functionally relevant cell populations
- More accurate modeling of tissue architecture and cellular ecosystems
- Deeper insights into disease mechanisms and therapeutic targets
Setting a New Standard in Spatial Biology
In spatial biology, chemistry enables detection, but segmentation defines biology. The ability to accurately assign molecules to the correct cell is essential for transforming spatial data into meaningful biological insight.
As spatial technologies continue to scale toward whole-transcriptome and multiomic measurements, the importance of accurate segmentation will only increase. High-plex data demands equally sophisticated computational approaches to ensure that biological conclusions remain valid and reproducible.
CosMx SMI represents a significant step forward—combining multi-focal imaging, multimodal data integration, and machine learning to deliver best-in-class segmentation performance. Ongoing innovations, including the integration of 3D segmentation models and purpose-trained segmentation models on diverse tissue types, promise to further enhance accuracy in densely packed and structurally complex tissues. The result is straightforward but critical: when the cell boundaries are correct, the biology becomes trustworthy.
References
- Petukhov et al., Nature Biotechnology (2022): Cell segmentation challenges in spatial transcriptomics. doi: 10.1038/s41587-021-01044.
- Liu et al., Biophys Rep (2022): Spatial transcriptomics: new dimension of understanding biological complexity. doi: 10.52601/bpr.2021.210037.
- Mitchel et al., Nature Genetics (2026): Impact and correction of segmentation errors in spatial transcriptomics. doi: 10.1038/s41588-025-02497-4.
- Defard et al., Genome Biology (2026): RNA2seg: a generalist model for cell segmentation in image-based spatial transcriptomics. doi: 10.1186/s13059-025-03908-9.
- Stringer et al., Nature Methods (2021): Cellpose: a generalist algorithm for cellular segmentation. doi: 10.1038/s41592-020-01018.
- Jin et al., Nature Communications (2025): Bering: joint cell segmentation and annotation for spatial transcriptomics with transferred graph embeddings. doi: 10.1101/2023.09.19.558548.
FAQs
What is cell segmentation in spatial transcriptomics?
Cell segmentation defines true cell boundaries to delineate molecular data to the correct functional cellular units (cells) in spatial transcriptomics.
Why is cell segmentation important?
Cell segmentation underpins all downstream analysis, directly impacting data accuracy and biological interpretation.
Why is cell segmentation challenging in complex tissues?
Cells exhibit heterogeneous shapes, variable sizes, and morphologies, making precise boundary definition challenging.
Dataset variability, including staining differences, imaging modalities, and tissue types further complicate accurate segmentation.
How are modern approaches improving cell segmentation accuracy?
Multimodal approaches combining RNA, nuclear, and membrane signals to better define true cell boundaries.
How does CosMx® SMI achieve best-in-class cell segmentation?
CosMx SMI integrates multimodal data and advanced machine learning to define true cellular boundaries. Bruker Spatial Biology further extends this capability with the 3D AI segmentation models by reconstructing cells across multiple focal planes. These models resolve overlapping cells and improve single-cell boundary definition in dense tissues, overcoming long-standing limitations of traditional segmentation approaches.